|
I am a Research Intern at the Centre for Brain Research, Indian Institute of Science (IISc), and completed my Master's degree in Data Science (2024-26) at Alliance University. Previously, I was a Summer ML Research Intern at the i2CS Research Group, IIIT Kottayam, where I worked with Dr. Kala S on multi-modal alignment of Vision Language Models for Plant Disease Detection. I graduated with a BSc in Mathematics and Statistics from Bengaluru City University. My research interests span Machine Learning, Computer Vision, NLP, and Generative Models, with applications to Neuroscience and Healthcare. Email / CV / LinkedIn / GitHub / Google Scholar Water Your Repo, Grow Your Greens! |

|
|
|
|
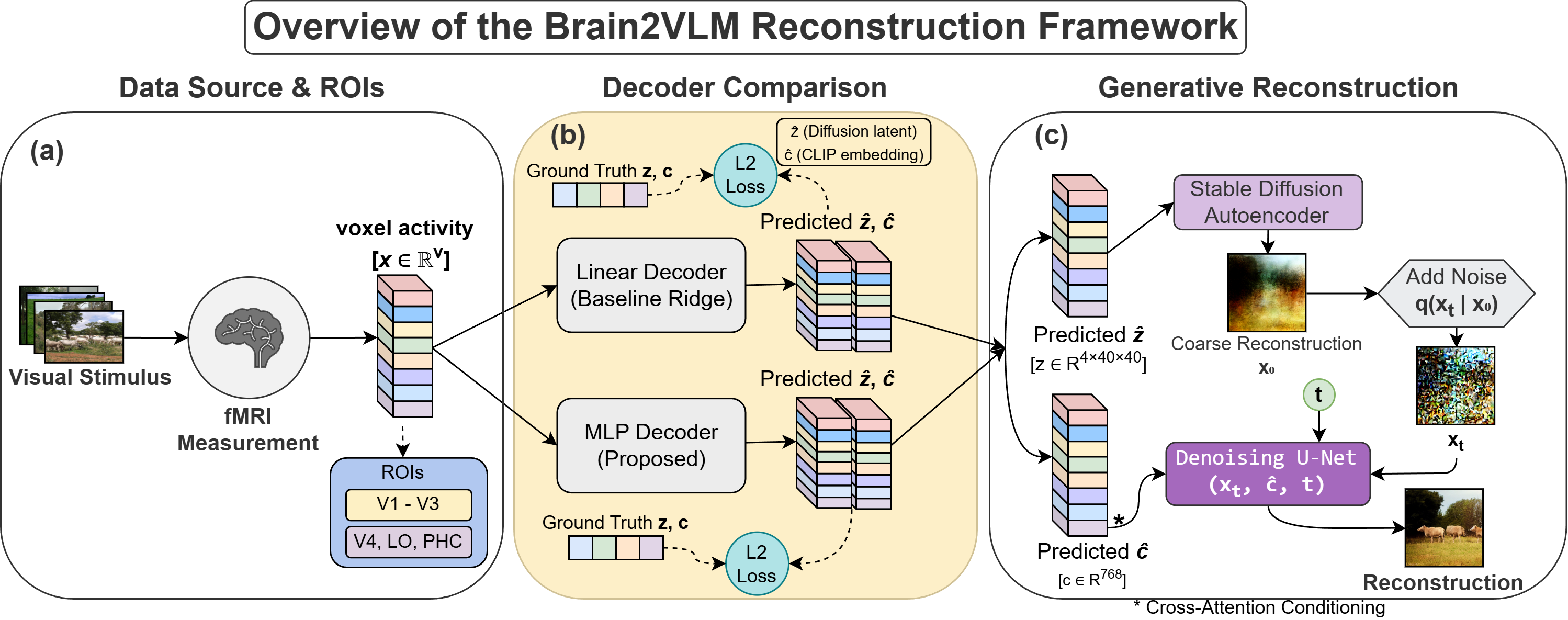
N A Adarsh Pritam , Jeba Shiney O, Sanyam Jain bioRxiv Preprint paper | code In this work, we study how visual stimuli can be recosntructed using fMRI signals by mapping neural activity to the latent spaces of pretrained diffusion-based vision-language models (e.g. LDM). Brain2VLM analyzes the structure of the brain-to-latent mapping rather than only focusing on improving reconstruction pipelines. We analyze how neural activity corresponds to structural diffusion latents and semantic embeddings, and how this varies across regions of the visual cortex. This provides a systematic view of brain-to-latent alignment for brain decoding. |
|
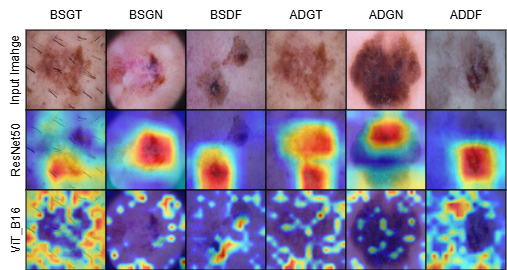
N A Adarsh Pritam , Jeba Shiney O, Sanyam Jain aRxiv Preprint paper | code In this work, we study how synthetic data can improve melanoma (skin cancer) diagnosis from dermoscopic images. SkinGenBench provides a controlled setup to analyze how generative model choice and preprocessing affect synthetic image augmentation in medical imaging. We compare GAN-based and diffusion-based approaches under different preprocessing conditions and evaluate their impact on image quality and downstream classification performance. This provides a systematic analysis of synthetic data augmentation for melanoma detection. |
|
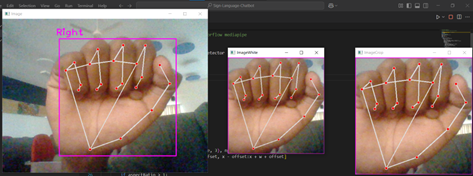
N A Adarsh Pritam , Asha Kurian IEEE - ETCC 2025 paper Developed a modular end-to-end pipeline for translating American Sign Language (ASL) fingerspelling from static images into English sentences, using Google Teachable Machine and a fine-tuned FLAN-T5 model for gloss-to-text generation. |
|
|

|
Principal Investigator (PI): Prof. Thomas Gregor Issac May 2026 - Present Researching the applications of Multimodal Machine Learning in Dementia Research for the early detection of Mild Cognitive Impairment using longitudinal clinical, cognitive, biochemical, and neuroimaging data from the ongoing flagship studies TLSA and SANSCOG. |

|
Supervisor(s): Prof. Jeba Shiney O Sep 2025 - May 2026 Conducted research as a part of master's thesis at Alliance University on applying machine learning to neuroscience and healthcare, including brain-to-image reconstruction from fMRI signals (Brain2VLM) and synthetic data augmentation for melanoma diagnosis (SkinGenBench). |

|
Supervisor(s): Dr. Kala S Provided deep learning edge to the group by developing a Vision-Language Model (VLM) for descriptive plant disease diagnosis by integrating a CLIP ViT-Large with an InternLM2-7B LLM. |
|
|
|
Proposed an interpretable statistical framework for modeling life expectancy using World Health Organization (WHO) data, used VIF-based multicollinearity resolution with domain-aware feature engineering and sequential feature selection. Developed and validated regression models within a reproducible pipeline, and deployed a Streamlit dashboard to analyze the influence of socioeconomic and healthcare variables, highlighting education, GDP, vaccination index, and BMI as key drivers. |
|
Updated Wei Yang's popular pytorch-classification repo by adding a unified training pipeline supporting 20+ architectures and automatic head adaptation for transfer learning. Also built an architecture-agnostic Grad-CAM system for clear interpretability across the entire model zoo. |
|
Implemented a LLaMA-inspired large language model from scratch in PyTorch, building core transformer components including multi-head attention, grouped-query attention (GQA), and rotary positional embeddings (RoPE). Analyzed the role of attention mechanisms and positional encoding in token representation and sequence modeling. |
|
|
|
For a complete overview of my open source contributions, check this GitHub search query. |
|
[merged] Contributed to improving documentation consistency and accessibility in the "Getting Started" guide by refining text clarity and metadata elements. [approved] Improved the model's positional embedding layer by refactoring duplicated logic to enhance code clarity and robustness. [merged] Improved code documentation by adding comprehensive docstrings to utility and helper functions. [merged] Improved an example script by removing dead code to enhance long-term code health and maintainability. [merged] Refactored a utility function for clarity, improving code readability and long-term maintainability. [merged] Resolved UnicodeDecodeError in a txt file-reader by specifying encoding for cross-platform compatibility. |
|
|
|
Master's in Data Science
(2024 - 2026) Alliance University, Bangalore, India Grade: 9.7/10 (Sem 4 SGPA) |
|
Bachelor of Science (BS) in Mathematics & Statistics
(2021 - 2024) St. Joseph's College (Autonomous), Bengaluru City University, Bangalore, India Grade: A |
|
|
| Base template taken from Nithish Kannen, thanks! |